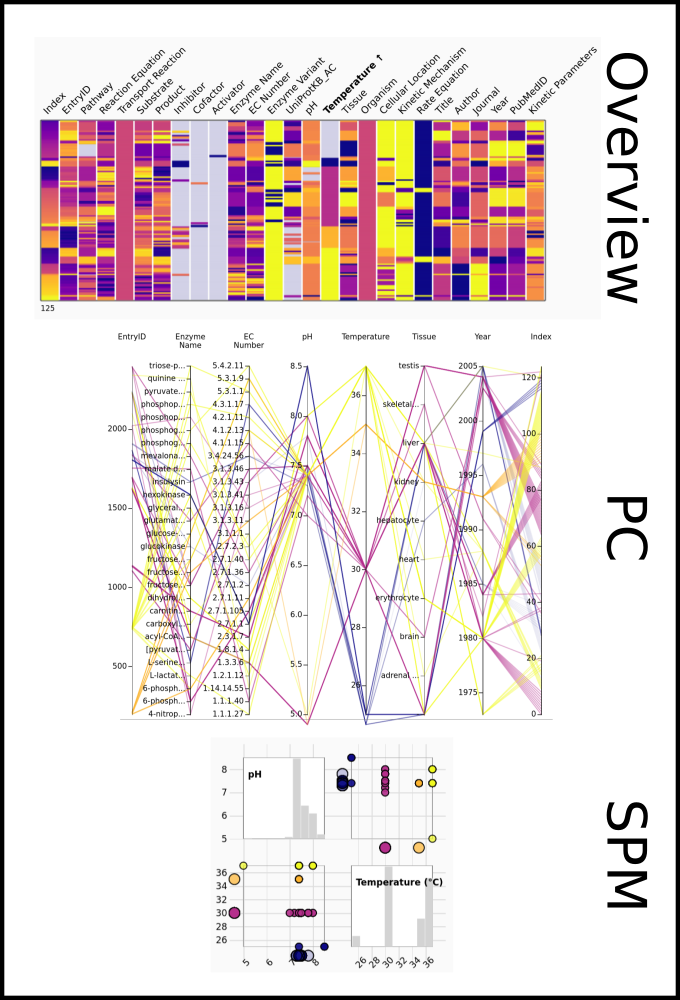
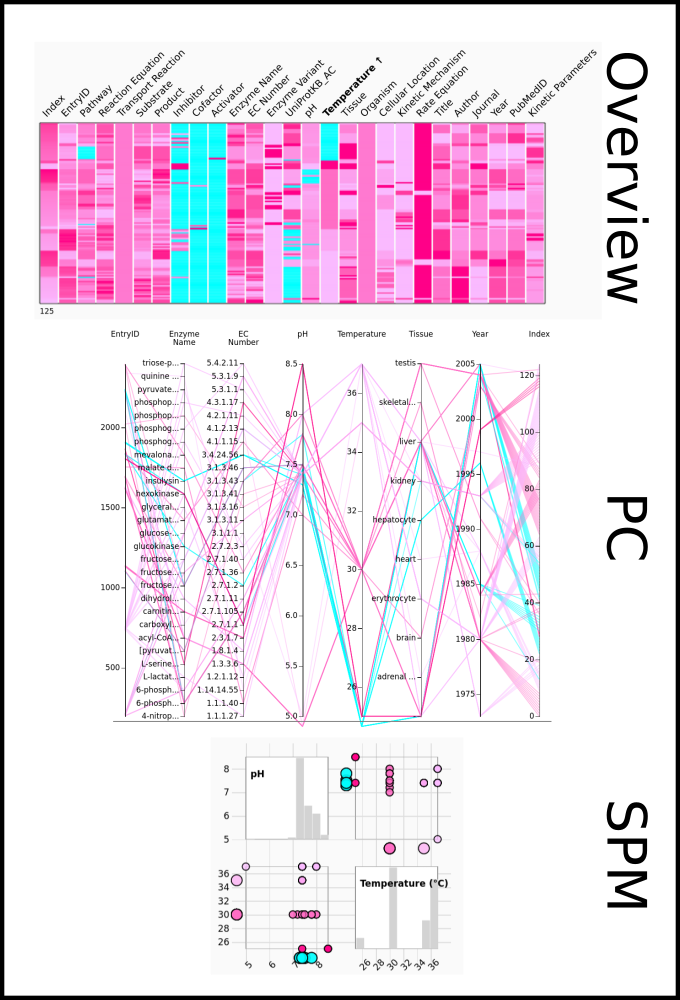
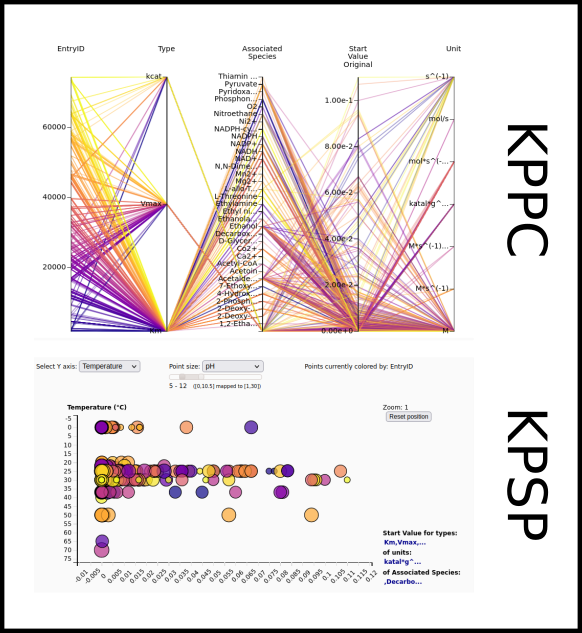
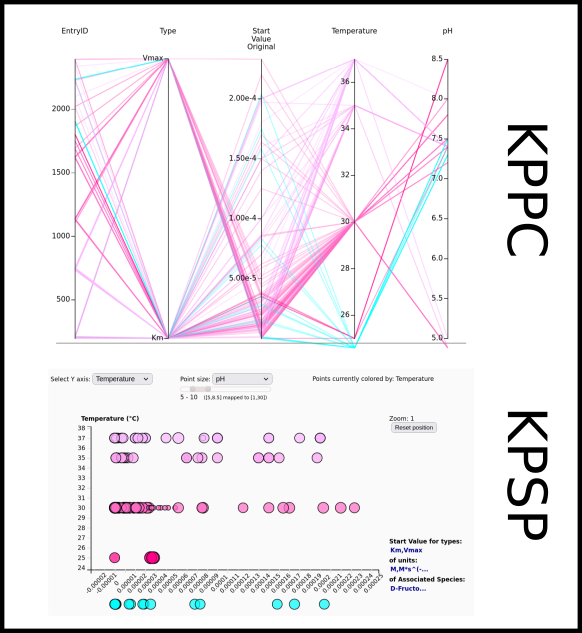
To improve SABIO-RK data interoperability, semantic markup was added to web pages as described and defined by
This structured information makes it easier to discover, collate and analyse our data.
|
61001
|
High-affinity, equilibrative nucleoside transporter of pig kidney cell line (PK-15)
|
|
61002
|
High-affinity, equilibrative nucleoside transporter of pig kidney cell line (PK-15)
|
|
61003
|
High-affinity, equilibrative nucleoside transporter of pig kidney cell line (PK-15)
|
|
61004
|
Structural basis and function of XRN2 binding by XTB domains
|
|
61005
|
Structural basis and function of XRN2 binding by XTB domains
|
|
61006
|
The generation of energy by the arginine dihydrolase pathway in Mycoplasma hominis 07.
|
|
61007
|
The generation of energy by the arginine dihydrolase pathway in Mycoplasma hominis 07.
|
|
61008
|
The generation of energy by the arginine dihydrolase pathway in Mycoplasma hominis 07.
|
|
61009
|
The generation of energy by the arginine dihydrolase pathway in Mycoplasma hominis 07.
|
|
61010
|
The generation of energy by the arginine dihydrolase pathway in Mycoplasma hominis 07.
|
|
61011
|
The generation of energy by the arginine dihydrolase pathway in Mycoplasma hominis 07.
|
|
61012
|
Structural and functional characterization of a 5,10-methenyltetrahydrofolate synthetase from Mycoplasma ...
|
|
61013
|
Structural and functional characterization of a 5,10-methenyltetrahydrofolate synthetase from Mycoplasma ...
|
|
61014
|
Structural and functional characterization of a 5,10-methenyltetrahydrofolate synthetase from Mycoplasma ...
|
|
61015
|
Structural and functional characterization of a 5,10-methenyltetrahydrofolate synthetase from Mycoplasma ...
|
|
61016
|
SIRT5 regulates the mitochondrial lysine succinylome and metabolic networks
|
|
61017
|
Discovery of a Glutamine Kinase Required for the Biosynthesis of the O-Methyl Phosphoramidate Modifications ...
|
|
61018
|
Discovery of a Glutamine Kinase Required for the Biosynthesis of the O-Methyl Phosphoramidate Modifications ...
|
|
61019
|
Biosynthesis of Nucleoside Diphosphoramidates in Campylobacter jejuni
|
|
61020
|
Biosynthesis of Nucleoside Diphosphoramidates in Campylobacter jejuni
|
|
61021
|
Biosynthesis of Nucleoside Diphosphoramidates in Campylobacter jejuni
|
|
61022
|
Monodehydroascorbate reductase mediates TNT toxicity in plants
|
|
61023
|
Monodehydroascorbate reductase mediates TNT toxicity in plants
|
|
61024
|
Monodehydroascorbate reductase mediates TNT toxicity in plants
|
|
61025
|
Activation of phenylalanine hydroxylase by phenylalanine does not require binding in the active site
|
|
61026
|
Hofmeister effects on activity and stability of alkaline phosphatase
|
|
61027
|
Hofmeister effects on activity and stability of alkaline phosphatase
|
|
61028
|
Hofmeister effects on activity and stability of alkaline phosphatase
|
|
61029
|
Hofmeister effects on activity and stability of alkaline phosphatase
|
|
61030
|
Hofmeister effects on activity and stability of alkaline phosphatase
|
|
61031
|
Hofmeister effects on activity and stability of alkaline phosphatase
|
|
61032
|
Hofmeister effects on activity and stability of alkaline phosphatase
|
|
61033
|
Hofmeister effects on activity and stability of alkaline phosphatase
|
|
61034
|
Hofmeister effects on activity and stability of alkaline phosphatase
|
|
61035
|
Hofmeister effects on activity and stability of alkaline phosphatase
|
|
61036
|
Hofmeister effects on activity and stability of alkaline phosphatase
|
|
61037
|
TrpB2 enzymes are O-phospho-l-serine dependent tryptophan synthases
|
|
61038
|
TrpB2 enzymes are O-phospho-l-serine dependent tryptophan synthases
|
|
61039
|
Mechanistic Studies of an Amine Oxidase Derived from d-Amino Acid Oxidase
|
|
61040
|
Mechanistic Studies of an Amine Oxidase Derived from d-Amino Acid Oxidase
|
|
61041
|
Mechanistic Studies of an Amine Oxidase Derived from d-Amino Acid Oxidase
|
|
61042
|
Identification and modulation of a growth hormone-binding protein in rainbow trout (Oncorhynchus mykiss) ...
|
|
61043
|
Identification and modulation of a growth hormone-binding protein in rainbow trout (Oncorhynchus mykiss) ...
|
|
61044
|
Identification and modulation of a growth hormone-binding protein in rainbow trout (Oncorhynchus mykiss) ...
|
|
61045
|
Identification and modulation of a growth hormone-binding protein in rainbow trout (Oncorhynchus mykiss) ...
|
|
61046
|
Mechanistic studies of the flavoprotein tryptophan 2-monooxygenase. 1. Kinetic mechanism
|
|
61047
|
Mechanistic studies of the flavoprotein tryptophan 2-monooxygenase. 1. Kinetic mechanism
|
|
61048
|
Mechanistic studies of the flavoprotein tryptophan 2-monooxygenase. 1. Kinetic mechanism
|
|
61049
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61050
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61051
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61052
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61053
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61054
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61055
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61056
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61057
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61058
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61059
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61060
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61061
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61062
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61063
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61064
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61065
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61066
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61067
|
Synthesis and enzymatic ketonization of the 5-(halo)-2-hydroxymuconates and ...
|
|
61068
|
Metal dependence and branched RNA cocrystal structures of the RNA lariat debranching enzyme Dbr1
|
|
61069
|
Human Neutrophil Elastase: Kinetics with Four Synthetic Substrates
|
|
61070
|
Human Neutrophil Elastase: Kinetics with Four Synthetic Substrates
|
|
61071
|
Human Neutrophil Elastase: Kinetics with Four Synthetic Substrates
|
|
61072
|
Human Neutrophil Elastase: Kinetics with Four Synthetic Substrates
|
|
61073
|
Human Myeloblastin: Kinetics with Four Synthetic Substrates
|
|
61074
|
Human Myeloblastin: Kinetics with Four Synthetic Substrates
|
|
61075
|
Human Myeloblastin: Kinetics with Four Synthetic Substrates
|
|
61076
|
Human Myeloblastin: Kinetics with Four Synthetic Substrates
|
|
61077
|
Human Myeloblastin: Interactions with Three Nonessential Activators
|
|
61078
|
Human Myeloblastin: Interactions with Three Nonessential Activators
|
|
61079
|
Human Myeloblastin: Interactions with Three Nonessential Activators
|
|
61080
|
Evolutionary diversification of protein-protein interactions by interface add-ons
|
|
61081
|
Evolutionary diversification of protein-protein interactions by interface add-ons
|
|
61082
|
Evolutionary diversification of protein-protein interactions by interface add-ons
|
|
61083
|
Evolutionary diversification of protein-protein interactions by interface add-ons
|
|
61084
|
Evolutionary diversification of protein-protein interactions by interface add-ons
|
|
61085
|
Structural and Enzymatic Basis for Oxamniquine Efficacy against Human Blood Fluke Parasites
|
|
61086
|
Structural and Enzymatic Basis for Oxamniquine Efficacy against Human Blood Fluke Parasites
|
|
61087
|
Structural and Enzymatic Basis for Oxamniquine Efficacy against Human Blood Fluke Parasites
|
|
61088
|
Structural and Enzymatic Basis for Oxamniquine Efficacy against Human Blood Fluke Parasites
|
|
61089
|
Structural and Enzymatic Basis for Oxamniquine Efficacy against Human Blood Fluke Parasites
|
|
61090
|
Structural and Enzymatic Basis for Oxamniquine Efficacy against Human Blood Fluke Parasites
|
|
61091
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61092
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61093
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61094
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61095
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61096
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61097
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61098
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61099
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61100
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61101
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61102
|
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe
|
|
61103
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61104
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61105
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61106
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61107
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61108
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61109
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61110
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61111
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61112
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61113
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61114
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61115
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61116
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61117
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61118
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61119
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61120
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61121
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61122
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61123
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61124
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61125
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61126
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61127
|
Evolutionarily Distinct BAHD N-Acyltransferases Are Responsible for Natural Variation of Aromatic Amine ...
|
|
61128
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61129
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61130
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61131
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61132
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61133
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61134
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61135
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61136
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61137
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61138
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61139
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61140
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61141
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61142
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61143
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61144
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61145
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61146
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61147
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61148
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61149
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61150
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61151
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61152
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61153
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61154
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61155
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61156
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61157
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61158
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61159
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61160
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61161
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61162
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61163
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61164
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61165
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61166
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61167
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61168
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61169
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61170
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61171
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61172
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61173
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61174
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61175
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61176
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61177
|
Reversible inhibition of sheep liver sorbitol dehydrogenase by thiol compounds.
|
|
61178
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61179
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61180
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61181
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61182
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61183
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61184
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61185
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61186
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61187
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61188
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61189
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61190
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61191
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61192
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61193
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61194
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61195
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61196
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61197
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61198
|
Kinetic mechanism of AKT/PKB enzyme family.
|
|
61199
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61200
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61201
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61202
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61203
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61204
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61205
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61206
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61207
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61208
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61209
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61210
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61211
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61212
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61213
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61214
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61215
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61216
|
ArcD1 and ArcD2 Arginine/Ornithine Exchangers Encoded in the Arginine Deiminase Pathway Gene Cluster of ...
|
|
61217
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61218
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61219
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61220
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61221
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61222
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61223
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61224
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61225
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61226
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61227
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61228
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61229
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61230
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61231
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61232
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61233
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61234
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61235
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61236
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61237
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61238
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61239
|
Catalysis of an Essential Step in Vitamin B2 Biosynthesis by a Consortium of Broad Spectrum Hydrolases
|
|
61240
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61241
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61242
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61243
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61244
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61245
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61246
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61247
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61248
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61249
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61250
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61251
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61252
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61253
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61254
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61255
|
Carbonyl reductase 1 is a predominant doxorubicin reductase in the human liver
|
|
61256
|
Cloning, sequencing, heterologous expression, purification, and characterization of ...
|
|
61257
|
Expression and characterization of RNase III and Era proteins. Products of the rnc operon of Escherichia coli.
|
|
61258
|
Expression and characterization of RNase III and Era proteins. Products of the rnc operon of Escherichia coli.
|
|
61259
|
Purification and properties of a malolactic enzyme from Leuconostoc oenos ATCC 23278
|
|
61260
|
Purification and properties of a malolactic enzyme from Leuconostoc oenos ATCC 23278
|
|
61261
|
Purification and properties of a malolactic enzyme from Leuconostoc oenos ATCC 23278
|
|
61262
|
Purification and properties of a malolactic enzyme from Leuconostoc oenos ATCC 23278
|
|
61263
|
Purification and properties of a malolactic enzyme from Leuconostoc oenos ATCC 23278
|
|
61264
|
Purification and properties of a malolactic enzyme from Leuconostoc oenos ATCC 23278
|
|
61265
|
Purification and properties of a malolactic enzyme from Leuconostoc oenos ATCC 23278
|
|
61266
|
Purification and properties of a malolactic enzyme from Leuconostoc oenos ATCC 23278
|
|
61267
|
Functional characterization of a novel ArgA from Mycobacterium tuberculosis.
|
|
61268
|
Functional characterization of a novel ArgA from Mycobacterium tuberculosis.
|
|
61269
|
Functional characterization of a novel ArgA from Mycobacterium tuberculosis.
|
|
61270
|
Functional characterization of a novel ArgA from Mycobacterium tuberculosis.
|
|
61271
|
Structure-based dissection of the active site chemistry of leukotriene A4 hydrolase: implications for M1 ...
|
|
61272
|
Structure-based dissection of the active site chemistry of leukotriene A4 hydrolase: implications for M1 ...
|
|
61273
|
Structure-based dissection of the active site chemistry of leukotriene A4 hydrolase: implications for M1 ...
|
|
61274
|
Structure-based dissection of the active site chemistry of leukotriene A4 hydrolase: implications for M1 ...
|
|
61275
|
Structure-based dissection of the active site chemistry of leukotriene A4 hydrolase: implications for M1 ...
|
|
61276
|
Structure-based dissection of the active site chemistry of leukotriene A4 hydrolase: implications for M1 ...
|
|
61277
|
Structure-based dissection of the active site chemistry of leukotriene A4 hydrolase: implications for M1 ...
|
|
61278
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61279
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61280
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61281
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61282
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61283
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61284
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61285
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61286
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61287
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61288
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61289
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61290
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61291
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61292
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61293
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61294
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61295
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61296
|
UDP-N-acetyl-α-D-galactosamine:polypeptide N-acetylgalactosaminyltransferases: completion of the family tree
|
|
61297
|
Nuclear-localized AtHSPR links abscisic acid-dependent salt tolerance and antioxidant defense in Arabidopsis.
|
|
61298
|
L-kynurenine: A fluorescent probe of serum albumins
|
|
61299
|
L-kynurenine: A fluorescent probe of serum albumins
|
|
61300
|
L-kynurenine: A fluorescent probe of serum albumins
|
|
61301
|
L-kynurenine: A fluorescent probe of serum albumins
|
|
61302
|
L-kynurenine: A fluorescent probe of serum albumins
|
|
61303
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61304
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61305
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61306
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61307
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61308
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61309
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61310
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61311
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61312
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61313
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61314
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61315
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61316
|
Flavonoids as inhibitors of human carbonyl reductase 1
|
|
61317
|
AVP2, a sequence-divergent, K(+)-insensitive H(+)-translocating inorganic pyrophosphatase from Arabidopsis
|
|
61318
|
AVP2, a sequence-divergent, K(+)-insensitive H(+)-translocating inorganic pyrophosphatase from Arabidopsis
|
|
61319
|
AVP2, a sequence-divergent, K(+)-insensitive H(+)-translocating inorganic pyrophosphatase from Arabidopsis
|
|
61320
|
AVP2, a sequence-divergent, K(+)-insensitive H(+)-translocating inorganic pyrophosphatase from Arabidopsis
|
|
61321
|
AVP2, a sequence-divergent, K(+)-insensitive H(+)-translocating inorganic pyrophosphatase from Arabidopsis
|
|
61322
|
AVP2, a sequence-divergent, K(+)-insensitive H(+)-translocating inorganic pyrophosphatase from Arabidopsis
|
|
61323
|
Octanoylation of early intermediates of mycobacterial methylglucose lipopolysaccharides
|
|
61324
|
Octanoylation of early intermediates of mycobacterial methylglucose lipopolysaccharides
|
|
61325
|
Octanoylation of early intermediates of mycobacterial methylglucose lipopolysaccharides
|
|
61326
|
Octanoylation of early intermediates of mycobacterial methylglucose lipopolysaccharides
|
|
61327
|
Octanoylation of early intermediates of mycobacterial methylglucose lipopolysaccharides
|
|
61328
|
Octanoylation of early intermediates of mycobacterial methylglucose lipopolysaccharides
|
|
61329
|
Octanoylation of early intermediates of mycobacterial methylglucose lipopolysaccharides
|
|
61330
|
Octanoylation of early intermediates of mycobacterial methylglucose lipopolysaccharides
|
|
61331
|
Octanoylation of early intermediates of mycobacterial methylglucose lipopolysaccharides
|
|
61332
|
Octanoylation of early intermediates of mycobacterial methylglucose lipopolysaccharides
|
|
61333
|
Octanoylation of early intermediates of mycobacterial methylglucose lipopolysaccharides
|
|
61334
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61335
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61336
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61337
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61338
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61339
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61340
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61341
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61342
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61343
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61344
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61345
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61346
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61347
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61348
|
Studies of 1-deoxy-D-fructose, 1-deoxy-D-glucitol, and 1-deoxy-D-minnitol as antimetabolites.
|
|
61349
|
Proteomic analysis and purification of an unusual germin-like protein with proteolytic activity in the latex ...
|
|
61350
|
Imidazole glycerol phosphate synthase from Thermotoga maritima. Quaternary structure, steady-state kinetics, ...
|
|
61351
|
Imidazole glycerol phosphate synthase from Thermotoga maritima. Quaternary structure, steady-state kinetics, ...
|
|
61352
|
Imidazole glycerol phosphate synthase from Thermotoga maritima. Quaternary structure, steady-state kinetics, ...
|
|
61353
|
Imidazole glycerol phosphate synthase from Thermotoga maritima. Quaternary structure, steady-state kinetics, ...
|
|
61354
|
Imidazole glycerol phosphate synthase from Thermotoga maritima. Quaternary structure, steady-state kinetics, ...
|
|
61355
|
Imidazole glycerol phosphate synthase from Thermotoga maritima. Quaternary structure, steady-state kinetics, ...
|
|
61356
|
Artificial domain duplication replicates evolutionary history of ketol-acid reductoisomerases
|
|
61357
|
Artificial domain duplication replicates evolutionary history of ketol-acid reductoisomerases
|
|
61358
|
Artificial domain duplication replicates evolutionary history of ketol-acid reductoisomerases
|
|
61359
|
Artificial domain duplication replicates evolutionary history of ketol-acid reductoisomerases
|
|
61360
|
Artificial domain duplication replicates evolutionary history of ketol-acid reductoisomerases
|
|
61361
|
Artificial domain duplication replicates evolutionary history of ketol-acid reductoisomerases
|
|
61362
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61363
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61364
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61365
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61366
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61367
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61368
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61369
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61370
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61371
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61372
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61373
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61374
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61375
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61376
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61377
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61378
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61379
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61380
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61381
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61382
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61383
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61384
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61385
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61386
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61387
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61388
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61389
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61390
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61391
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61392
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61393
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61394
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61395
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61396
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61397
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61398
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61399
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61400
|
A study on reducing substrates of manganese-oxidizing peroxidases from Pleurotus eryngii and Bjerkandera ...
|
|
61401
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61402
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61403
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61404
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61405
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61406
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61407
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61408
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61409
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61410
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61411
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61412
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61413
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61414
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61415
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61416
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61417
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61418
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61419
|
Different fungal manganese-oxidizing peroxidases: a comparison between Bjerkandera sp. and Phanerochaete ...
|
|
61420
|
Mutation of a nitrate transporter, AtNRT1:4, results in a reduced petiole nitrate content and altered leaf ...
|
|
61421
|
The Thermoplasma acidophilum Lon protease has a Ser-Lys dyad active site
|
|
61422
|
The Thermoplasma acidophilum Lon protease has a Ser-Lys dyad active site
|
|
61423
|
The Thermoplasma acidophilum Lon protease has a Ser-Lys dyad active site
|
|
61424
|
The Thermoplasma acidophilum Lon protease has a Ser-Lys dyad active site
|
|
61425
|
The Thermoplasma acidophilum Lon protease has a Ser-Lys dyad active site
|
|
61426
|
The Thermoplasma acidophilum Lon protease has a Ser-Lys dyad active site
|
|
61427
|
NADP-dependent isocitrate dehydrogenase from the halophilic archaeon Haloferax volcanii: cloning, sequence ...
|
|
61428
|
Isocitrate dehydrogenases from Haloferax volcanii and Sulfolobus solfataricus: enzyme purification, ...
|
|
61429
|
Isocitrate dehydrogenases from Haloferax volcanii and Sulfolobus solfataricus: enzyme purification, ...
|
|
61430
|
Isocitrate dehydrogenases from Haloferax volcanii and Sulfolobus solfataricus: enzyme purification, ...
|
|
61431
|
NMR study of manganese(II) binding by a new versatile peroxidase from the white-rot fungus Pleurotus eryngii
|
|
61432
|
NMR study of manganese(II) binding by a new versatile peroxidase from the white-rot fungus Pleurotus eryngii
|
|
61433
|
NMR study of manganese(II) binding by a new versatile peroxidase from the white-rot fungus Pleurotus eryngii
|
|
61434
|
NMR study of manganese(II) binding by a new versatile peroxidase from the white-rot fungus Pleurotus eryngii
|
|
61435
|
NMR study of manganese(II) binding by a new versatile peroxidase from the white-rot fungus Pleurotus eryngii
|
|
61436
|
NMR study of manganese(II) binding by a new versatile peroxidase from the white-rot fungus Pleurotus eryngii
|
|
61437
|
NMR study of manganese(II) binding by a new versatile peroxidase from the white-rot fungus Pleurotus eryngii
|
|
61438
|
NMR study of manganese(II) binding by a new versatile peroxidase from the white-rot fungus Pleurotus eryngii
|
|
61439
|
Crystal structure of human wildtype and S581L-mutant glycyl-tRNA synthetase, an enzyme underlying distal ...
|
|
61440
|
Crystal structure of human wildtype and S581L-mutant glycyl-tRNA synthetase, an enzyme underlying distal ...
|
|
61441
|
Catalytic mechanism revealed by the crystal structure of undecaprenyl pyrophosphate synthase in complex with ...
|
|
61442
|
Catalytic mechanism revealed by the crystal structure of undecaprenyl pyrophosphate synthase in complex with ...
|
|
61443
|
Catalytic mechanism revealed by the crystal structure of undecaprenyl pyrophosphate synthase in complex with ...
|
|
61444
|
Catalytic mechanism revealed by the crystal structure of undecaprenyl pyrophosphate synthase in complex with ...
|
|
61445
|
Catalytic mechanism revealed by the crystal structure of undecaprenyl pyrophosphate synthase in complex with ...
|
|
61446
|
Catalytic mechanism revealed by the crystal structure of undecaprenyl pyrophosphate synthase in complex with ...
|
|
61447
|
Catalytic mechanism revealed by the crystal structure of undecaprenyl pyrophosphate synthase in complex with ...
|
|
61448
|
Catalytic mechanism revealed by the crystal structure of undecaprenyl pyrophosphate synthase in complex with ...
|
|
61449
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61450
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61451
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61452
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61453
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61454
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61455
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61456
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61457
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61458
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61459
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61460
|
Functional characterization of two novel mammalian electrogenic proton-dependent amino acid cotransporters
|
|
61461
|
Biochemical characterization of proline racemases from the human protozoan parasite Trypanosoma cruzi and ...
|
|
61462
|
Biochemical characterization of proline racemases from the human protozoan parasite Trypanosoma cruzi and ...
|
|
61463
|
Biochemical characterization of proline racemases from the human protozoan parasite Trypanosoma cruzi and ...
|
|
61464
|
Biochemical characterization of proline racemases from the human protozoan parasite Trypanosoma cruzi and ...
|
|
61465
|
Comparison of the three rat GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferases FTA, FTB and FTC
|
|
61466
|
Comparison of the three rat GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferases FTA, FTB and FTC
|
|
61467
|
Comparison of the three rat GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferases FTA, FTB and FTC
|
|
61468
|
Comparison of the three rat GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferases FTA, FTB and FTC
|
|
61469
|
Comparison of the three rat GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferases FTA, FTB and FTC
|
|
61470
|
Comparison of the three rat GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferases FTA, FTB and FTC
|
|
61471
|
Comparison of the three rat GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferases FTA, FTB and FTC
|
|
61472
|
Comparison of the three rat GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferases FTA, FTB and FTC
|
|
61473
|
Comparison of the three rat GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferases FTA, FTB and FTC
|
|
61474
|
Comparison of the three rat GDP-L-fucose:beta-D-galactoside 2-alpha-L-fucosyltransferases FTA, FTB and FTC
|
|
61475
|
Molecular cloning, chromosomal assignment and tissue-specific expression of a murine ...
|
|
61476
|
Molecular cloning, chromosomal assignment and tissue-specific expression of a murine ...
|
|
61477
|
Molecular cloning, chromosomal assignment and tissue-specific expression of a murine ...
|
|
61478
|
Molecular cloning, chromosomal assignment and tissue-specific expression of a murine ...
|
|
61479
|
Molecular cloning, chromosomal assignment and tissue-specific expression of a murine ...
|
|
61480
|
Molecular cloning, chromosomal assignment and tissue-specific expression of a murine ...
|
|
61481
|
Molecular cloning, chromosomal assignment and tissue-specific expression of a murine ...
|
|
61482
|
Molecular cloning, chromosomal assignment and tissue-specific expression of a murine ...
|
|
61483
|
Molecular cloning, chromosomal assignment and tissue-specific expression of a murine ...
|
|
61484
|
Molecular cloning, chromosomal assignment and tissue-specific expression of a murine ...
|
|
61485
|
Molecular cloning, chromosomal assignment and tissue-specific expression of a murine ...
|
|
61486
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61487
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61488
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61489
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61490
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61491
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61492
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61493
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61494
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61495
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61496
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61497
|
Design and synthesis of 1,3-diarylurea derivatives as selective cyclooxygenase (COX-2) inhibitors
|
|
61498
|
Two mechanisms for mannose-binding protein modulation of the activity of its associated serine proteases
|
|
61499
|
Two mechanisms for mannose-binding protein modulation of the activity of its associated serine proteases
|
|
61500
|
Two mechanisms for mannose-binding protein modulation of the activity of its associated serine proteases
|
|
61501
|
Two mechanisms for mannose-binding protein modulation of the activity of its associated serine proteases
|
|
61502
|
Two mechanisms for mannose-binding protein modulation of the activity of its associated serine proteases
|
|
61503
|
Two mechanisms for mannose-binding protein modulation of the activity of its associated serine proteases
|
|
61504
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61505
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61506
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61507
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61508
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61509
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61510
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61511
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61512
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61513
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61514
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61515
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61516
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61517
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61518
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61519
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61520
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61521
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61522
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61523
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61524
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61525
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61526
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61527
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61528
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61529
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61530
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61531
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61532
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61533
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61534
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61535
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61536
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61537
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61538
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61539
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61540
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61541
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61542
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61543
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61544
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61545
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61546
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61547
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61548
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61549
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61550
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61551
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61552
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61553
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61554
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61555
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61556
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61557
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61558
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61559
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61560
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61561
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61562
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61563
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61564
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61565
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61566
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61567
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61568
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61569
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61570
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61571
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61572
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61573
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61574
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61575
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61576
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61577
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61578
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61579
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61580
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61581
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61582
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61583
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61584
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61585
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61586
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61587
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61588
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61589
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61590
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61591
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61592
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61593
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61594
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61595
|
Glucokinase and hexokinase expression and activities in rainbow trout tissues: changes with food deprivation ...
|
|
61596
|
Purification and properties of nitroalkane oxidase from Fusarium oxysporum.
|
|
61597
|
Purification and properties of nitroalkane oxidase from Fusarium oxysporum.
|
|
61598
|
Purification and properties of nitroalkane oxidase from Fusarium oxysporum.
|
|
61599
|
Purification and properties of nitroalkane oxidase from Fusarium oxysporum.
|
|
61600
|
Purification and properties of nitroalkane oxidase from Fusarium oxysporum.
|
|
61601
|
Purification and properties of nitroalkane oxidase from Fusarium oxysporum.
|
|
61602
|
Purification of the o-dianisidine peroxidase from Escherichia coli B. Physicochemical characterization and ...
|
|
61603
|
Purification of the o-dianisidine peroxidase from Escherichia coli B. Physicochemical characterization and ...
|
|
61604
|
Purification and properties of Bacillus subtilis inositol dehydrogenase.
|
|
61605
|
Purification and properties of Bacillus subtilis inositol dehydrogenase.
|
|
61606
|
Purification and properties of Bacillus subtilis inositol dehydrogenase.
|
|
61607
|
Purification and properties of Bacillus subtilis inositol dehydrogenase.
|
|
61608
|
Purification and properties of Bacillus subtilis inositol dehydrogenase.
|
|
61609
|
Purification and properties of Bacillus subtilis inositol dehydrogenase.
|
|
61610
|
Purification and properties of Bacillus subtilis inositol dehydrogenase.
|
|
61611
|
Purification and properties of Bacillus subtilis inositol dehydrogenase.
|
|
61612
|
Purification and properties of Bacillus subtilis inositol dehydrogenase.
|
|
61613
|
Enzymes of glucose and methanol metabolism in the actinomycete Amycolatopsis methanolica
|
|
61614
|
Enzymes of glucose and methanol metabolism in the actinomycete Amycolatopsis methanolica
|
|
61615
|
Enzymes of glucose and methanol metabolism in the actinomycete Amycolatopsis methanolica
|
|
61616
|
Enzymes of glucose and methanol metabolism in the actinomycete Amycolatopsis methanolica
|
|
61617
|
Enzymes of glucose and methanol metabolism in the actinomycete Amycolatopsis methanolica
|
|
61618
|
Enzymes of glucose and methanol metabolism in the actinomycete Amycolatopsis methanolica
|
|
61619
|
Enzymes of glucose and methanol metabolism in the actinomycete Amycolatopsis methanolica
|
|
61620
|
Enzymes of glucose and methanol metabolism in the actinomycete Amycolatopsis methanolica
|
|
61621
|
Enzymes of glucose and methanol metabolism in the actinomycete Amycolatopsis methanolica
|
|
61622
|
Enzymes of glucose and methanol metabolism in the actinomycete Amycolatopsis methanolica
|
|
61623
|
Enzymes of glucose and methanol metabolism in the actinomycete Amycolatopsis methanolica
|
|
61624
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61625
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61626
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61627
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61628
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61629
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61630
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61631
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61632
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61633
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61634
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61635
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61636
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61637
|
Cloning and characterization of a mammalian lithium-sensitive bisphosphate 3'-nucleotidase inhibited by ...
|
|
61638
|
Molecular cloning and functional characterization of a novel mammalian sphingosine kinase type 2 isoform
|
|
61639
|
Molecular cloning and functional characterization of a novel mammalian sphingosine kinase type 2 isoform
|
|
61640
|
Molecular cloning and functional characterization of a novel mammalian sphingosine kinase type 2 isoform
|
|
61641
|
Molecular cloning and functional characterization of a novel mammalian sphingosine kinase type 2 isoform
|
|
61642
|
Molecular cloning and functional characterization of a novel mammalian sphingosine kinase type 2 isoform
|
|
61643
|
Molecular cloning and functional characterization of a novel mammalian sphingosine kinase type 2 isoform
|
|
61644
|
Molecular cloning and functional characterization of a novel mammalian sphingosine kinase type 2 isoform
|
|
61645
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61646
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61647
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61648
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61649
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61650
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61651
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61652
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61653
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61654
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61655
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61656
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61657
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61658
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61659
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61660
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61661
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61662
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61663
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61664
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61665
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61666
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61667
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61668
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61669
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61670
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61671
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61672
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61673
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61674
|
Interaction of estrogen mimics, singly and in combination, with plasma sex steroid-binding proteins in rainbow ...
|
|
61675
|
Enzymological properties of pantothenate synthetase from Escherichia coli B.
|
|
61676
|
Enzymological properties of pantothenate synthetase from Escherichia coli B.
|
|
61677
|
Enzymological properties of pantothenate synthetase from Escherichia coli B.
|
|
61678
|
Enzymological properties of pantothenate synthetase from Escherichia coli B.
|
|
61679
|
Enzymological properties of pantothenate synthetase from Escherichia coli B.
|
|
61680
|
Three different systems participate in L-cystine uptake in Bacillus subtilis
|
|
61681
|
Three different systems participate in L-cystine uptake in Bacillus subtilis
|
|
61682
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61683
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61684
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61685
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61686
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61687
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61688
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61689
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61690
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61691
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61692
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61693
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61694
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61695
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61696
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61697
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61698
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61699
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61700
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61701
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61702
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61703
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61704
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61705
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61706
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61707
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61708
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61709
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61710
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61711
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61712
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61713
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61714
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61715
|
Biochemical properties of nematode O-acetylserine(thiol)lyase paralogs imply their distinct roles in hydrogen ...
|
|
61716
|
Elucidation of active site residues of Arabidopsis thaliana flavonol synthase provides a molecular platform ...
|
|
61717
|
Elucidation of active site residues of Arabidopsis thaliana flavonol synthase provides a molecular platform ...
|
|
61718
|
Elucidation of active site residues of Arabidopsis thaliana flavonol synthase provides a molecular platform ...
|
|
61719
|
Elucidation of active site residues of Arabidopsis thaliana flavonol synthase provides a molecular platform ...
|
|
61720
|
Elucidation of active site residues of Arabidopsis thaliana flavonol synthase provides a molecular platform ...
|
|
61721
|
Elucidation of active site residues of Arabidopsis thaliana flavonol synthase provides a molecular platform ...
|
|
61722
|
Functional characterization of tyrosine transport in fibroblast cells from healthy controls
|
|
61723
|
Functional characterization of tyrosine transport in fibroblast cells from healthy controls
|
|
61724
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61725
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61726
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61727
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61728
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61729
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61730
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61731
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61732
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61733
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61734
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61735
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61736
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61737
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61738
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61739
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61740
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61741
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61742
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61743
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61744
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61745
|
Differential regulation of opposing RelMtb activities by the aminoacylation state of a ...
|
|
61746
|
Characterization of an acyltransferase capable of synthesizing benzylbenzoate and other volatile esters in ...
|
|
61747
|
Characterization of an acyltransferase capable of synthesizing benzylbenzoate and other volatile esters in ...
|
|
61748
|
Characterization of an acyltransferase capable of synthesizing benzylbenzoate and other volatile esters in ...
|
|
61749
|
Characterization of an acyltransferase capable of synthesizing benzylbenzoate and other volatile esters in ...
|
|
61750
|
Characterization of an acyltransferase capable of synthesizing benzylbenzoate and other volatile esters in ...
|
|
61751
|
Characterization of an acyltransferase capable of synthesizing benzylbenzoate and other volatile esters in ...
|
|
61752
|
Characterization of an acyltransferase capable of synthesizing benzylbenzoate and other volatile esters in ...
|
|
61753
|
Characterization of an acyltransferase capable of synthesizing benzylbenzoate and other volatile esters in ...
|
|
61754
|
Characterization of an acyltransferase capable of synthesizing benzylbenzoate and other volatile esters in ...
|
|
61755
|
Avian multiple inositol polyphosphate phosphatase is an active phytase that can be engineered to help ...
|
|
61756
|
Avian multiple inositol polyphosphate phosphatase is an active phytase that can be engineered to help ...
|
|
61757
|
Avian multiple inositol polyphosphate phosphatase is an active phytase that can be engineered to help ...
|
|
61758
|
Avian multiple inositol polyphosphate phosphatase is an active phytase that can be engineered to help ...
|
|
61759
|
Avian multiple inositol polyphosphate phosphatase is an active phytase that can be engineered to help ...
|
|
61760
|
Avian multiple inositol polyphosphate phosphatase is an active phytase that can be engineered to help ...
|
|
61761
|
Characterization of the transport properties of human multidrug resistance protein 7 (MRP7, ABCC10)
|
|
61762
|
Characterization of the transport properties of human multidrug resistance protein 7 (MRP7, ABCC10)
|
|
61763
|
Characterization of the transport properties of human multidrug resistance protein 7 (MRP7, ABCC10)
|
|
61764
|
Characterization of the transport properties of human multidrug resistance protein 7 (MRP7, ABCC10)
|
|
61765
|
Characterization of the transport properties of human multidrug resistance protein 7 (MRP7, ABCC10)
|
|
61766
|
Transformation of isopropylamine to L-alaninol by Pseudomonas sp. strain KIE171 involves N-glutamylated ...
|
|
61767
|
Transformation of isopropylamine to L-alaninol by Pseudomonas sp. strain KIE171 involves N-glutamylated ...
|
|
61768
|
Factors affecting the level and activity of pyruvate kinase from Coprinus lagopus sensu Buller.
|
|
61769
|
Factors affecting the level and activity of pyruvate kinase from Coprinus lagopus sensu Buller.
|
|
61770
|
Factors affecting the level and activity of pyruvate kinase from Coprinus lagopus sensu Buller.
|
|
61771
|
Factors affecting the level and activity of pyruvate kinase from Coprinus lagopus sensu Buller.
|
|
61772
|
Factors affecting the level and activity of pyruvate kinase from Coprinus lagopus sensu Buller.
|
|
61773
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61774
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61775
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61776
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61777
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61778
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61779
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61780
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61781
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61782
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61783
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61784
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61785
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61786
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61787
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61788
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61789
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61790
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61791
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61792
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61793
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61794
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61795
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61796
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61797
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61798
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61799
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61800
|
Role of RecJ-like protein with 5'-3' exonuclease activity in oligo(deoxy)nucleotide degradation.
|
|
61801
|
Mouse MTH2 protein which prevents mutations caused by 8-oxoguanine nucleotides
|
|
61802
|
Mouse MTH2 protein which prevents mutations caused by 8-oxoguanine nucleotides
|
|
61803
|
Inhibition of aldose reductase by dietary antioxidant curcumin: mechanism of inhibition, specificity and ...
|
|
61804
|
Inhibition of aldose reductase by dietary antioxidant curcumin: mechanism of inhibition, specificity and ...
|
|
61805
|
Inhibition of aldose reductase by dietary antioxidant curcumin: mechanism of inhibition, specificity and ...
|
|
61806
|
Inhibition of aldose reductase by dietary antioxidant curcumin: mechanism of inhibition, specificity and ...
|
|
61807
|
Inhibition of aldose reductase by dietary antioxidant curcumin: mechanism of inhibition, specificity and ...
|
|
61808
|
Inhibition of aldose reductase by dietary antioxidant curcumin: mechanism of inhibition, specificity and ...
|
|
61809
|
Inhibition of aldose reductase by dietary antioxidant curcumin: mechanism of inhibition, specificity and ...
|
|
61810
|
Expression of soluble and catalytically active plant (monocot) beta-glucosidases in E. coli.
|
|
61811
|
Expression of soluble and catalytically active plant (monocot) beta-glucosidases in E. coli.
|
|
61812
|
Expression of soluble and catalytically active plant (monocot) beta-glucosidases in E. coli.
|
|
61813
|
Expression of soluble and catalytically active plant (monocot) beta-glucosidases in E. coli.
|
|
61814
|
Expression of soluble and catalytically active plant (monocot) beta-glucosidases in E. coli.
|
|
61815
|
Expression of soluble and catalytically active plant (monocot) beta-glucosidases in E. coli.
|
|
61816
|
Expression of soluble and catalytically active plant (monocot) beta-glucosidases in E. coli.
|
|
61817
|
Expression of soluble and catalytically active plant (monocot) beta-glucosidases in E. coli.
|
|
61818
|
Mechanistic studies of two dioxygenases in the methionine salvage pathway of Klebsiella pneumoniae.
|
|
61819
|
Mechanistic studies of two dioxygenases in the methionine salvage pathway of Klebsiella pneumoniae.
|
|
61820
|
Molecular cloning and expression of a functional snake venom serine proteinase, with platelet aggregating ...
|
|
61821
|
Molecular cloning and expression of a functional snake venom serine proteinase, with platelet aggregating ...
|
|
61822
|
Molecular cloning and expression of a functional snake venom serine proteinase, with platelet aggregating ...
|
|
61823
|
Molecular cloning and expression of a functional snake venom serine proteinase, with platelet aggregating ...
|
|
61824
|
2-Methylisocitrate lyases from the bacterium Escherichia coli and the filamentous fungus Aspergillus nidulans: ...
|
|
61825
|
2-Methylisocitrate lyases from the bacterium Escherichia coli and the filamentous fungus Aspergillus nidulans: ...
|
|
61826
|
2-Methylisocitrate lyases from the bacterium Escherichia coli and the filamentous fungus Aspergillus nidulans: ...
|
|
61827
|
2-Methylisocitrate lyases from the bacterium Escherichia coli and the filamentous fungus Aspergillus nidulans: ...
|
|
61828
|
Generation and phenotypic characterization of Aspergillus nidulans methylisocitrate lyase deletion mutants: ...
|
|
61829
|
Generation and phenotypic characterization of Aspergillus nidulans methylisocitrate lyase deletion mutants: ...
|
|
61830
|
Generation and phenotypic characterization of Aspergillus nidulans methylisocitrate lyase deletion mutants: ...
|
|
61831
|
Generation and phenotypic characterization of Aspergillus nidulans methylisocitrate lyase deletion mutants: ...
|
|
61832
|
Generation and phenotypic characterization of Aspergillus nidulans methylisocitrate lyase deletion mutants: ...
|
|
61833
|
Generation and phenotypic characterization of Aspergillus nidulans methylisocitrate lyase deletion mutants: ...
|
|
61834
|
Functional Annotation of LigU as a 1,3-Allylic Isomerase during the Degradation of Lignin in the ...
|
|
61835
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61836
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61837
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61838
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61839
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61840
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61841
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61842
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61843
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61844
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61845
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61846
|
Brain-specific tryptophan hydroxylase 2 (TPH2): a functional Pro206Ser substitution and variation in the ...
|
|
61847
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61848
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61849
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61850
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61851
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61852
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61853
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61854
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61855
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61856
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61857
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61858
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61859
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61860
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61861
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61862
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61863
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61864
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61865
|
Cloning and characterization of a functional human gamma-aminobutyric acid (GABA) transporter, human GAT-2.
|
|
61866
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61867
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61868
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61869
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61870
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61871
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61872
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61873
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61874
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61875
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61876
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61877
|
The PII signal transduction protein of Arabidopsis thaliana forms an arginine-regulated complex with plastid ...
|
|
61878
|
Factorizing the role of a critical leucine residue in the binding of substrate to human 20alpha-hydroxysteroid ...
|
|
61879
|
Factorizing the role of a critical leucine residue in the binding of substrate to human 20alpha-hydroxysteroid ...
|
|
61880
|
Factorizing the role of a critical leucine residue in the binding of substrate to human 20alpha-hydroxysteroid ...
|
|
61881
|
Factorizing the role of a critical leucine residue in the binding of substrate to human 20alpha-hydroxysteroid ...
|
|
61882
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61883
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61884
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61885
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61886
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61887
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61888
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61889
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61890
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61891
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61892
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61893
|
Design, synthesis and biological evaluation of new 2,3-diarylquinoline derivatives as selective ...
|
|
61894
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61895
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61896
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61897
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61898
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61899
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61900
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61901
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61902
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61903
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61904
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61905
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61906
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61907
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61908
|
Design, synthesis, and biological evaluation of ketoprofen analogs as potent cyclooxygenase-2 inhibitors.
|
|
61909
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61910
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61911
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61912
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61913
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61914
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61915
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61916
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61917
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61918
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61919
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61920
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61921
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61922
|
Design and synthesis of new 1,3-benzthiazinan-4-one derivatives as selective cyclooxygenase (COX-2) ...
|
|
61923
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61924
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61925
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61926
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61927
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61928
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61929
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61930
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61931
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61932
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61933
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61934
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61935
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61936
|
Synthesis and biological evaluation of new 4-carboxyl quinoline derivatives as cyclooxygenase-2 inhibitors.
|
|
61937
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61938
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61939
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61940
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61941
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61942
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61943
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61944
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61945
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61946
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61947
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61948
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61949
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61950
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61951
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61952
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61953
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61954
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61955
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61956
|
Site-directed mutagenesis and halophilicity of dihydrolipoamide dehydrogenase from the halophilic archaeon, ...
|
|
61957
|
Glutathione S-transferase interacting with far-red insensitive 219 is involved in phytochrome A-mediated ...
|
|
61958
|
Glutathione S-transferase interacting with far-red insensitive 219 is involved in phytochrome A-mediated ...
|
|
61959
|
Characterization of human lysophospholipid acyltransferase 3.
|
|
61960
|
Characterization of human lysophospholipid acyltransferase 3.
|
|
61961
|
Characterization of human lysophospholipid acyltransferase 3.
|
|
61962
|
Characterization of human lysophospholipid acyltransferase 3.
|
|
61963
|
Characterization of human lysophospholipid acyltransferase 3.
|
|
61964
|
Characterization of human lysophospholipid acyltransferase 3.
|
|
61965
|
Characterization of human lysophospholipid acyltransferase 3.
|
|
61966
|
Characterization of human lysophospholipid acyltransferase 3.
|
|
61967
|
Characterization of human lysophospholipid acyltransferase 3.
|
|
61968
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61969
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61970
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61971
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61972
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61973
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61974
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61975
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61976
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61977
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61978
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61979
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61980
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61981
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61982
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61983
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61984
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61985
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61986
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61987
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61988
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61989
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61990
|
Catalysis by the second class of tRNA(m1G37) methyl transferase requires a conserved proline.
|
|
61991
|
The non-photosynthetic phosphoenolpyruvate carboxylases of the C4 dicot Flaveria trinervia -- implications for ...
|
|
61992
|
The non-photosynthetic phosphoenolpyruvate carboxylases of the C4 dicot Flaveria trinervia -- implications for ...
|
|
61993
|
The non-photosynthetic phosphoenolpyruvate carboxylases of the C4 dicot Flaveria trinervia -- implications for ...
|
|
61994
|
The non-photosynthetic phosphoenolpyruvate carboxylases of the C4 dicot Flaveria trinervia -- implications for ...
|
|
61995
|
The non-photosynthetic phosphoenolpyruvate carboxylases of the C4 dicot Flaveria trinervia -- implications for ...
|
|
61996
|
The non-photosynthetic phosphoenolpyruvate carboxylases of the C4 dicot Flaveria trinervia -- implications for ...
|
|
61997
|
The non-photosynthetic phosphoenolpyruvate carboxylases of the C4 dicot Flaveria trinervia -- implications for ...
|
|
61998
|
The non-photosynthetic phosphoenolpyruvate carboxylases of the C4 dicot Flaveria trinervia -- implications for ...
|
|
61999
|
The non-photosynthetic phosphoenolpyruvate carboxylases of the C4 dicot Flaveria trinervia -- implications for ...
|
|
62000
|
The non-photosynthetic phosphoenolpyruvate carboxylases of the C4 dicot Flaveria trinervia -- implications for ...
|





 If the current search query is a refinement of the previous query "Use Previous Query (Go Back)" button can be used to perform the previous query again. Only one automatic step back is possible, but the query can be manually manipulated if desired.
If the current search query is a refinement of the previous query "Use Previous Query (Go Back)" button can be used to perform the previous query again. Only one automatic step back is possible, but the query can be manually manipulated if desired.